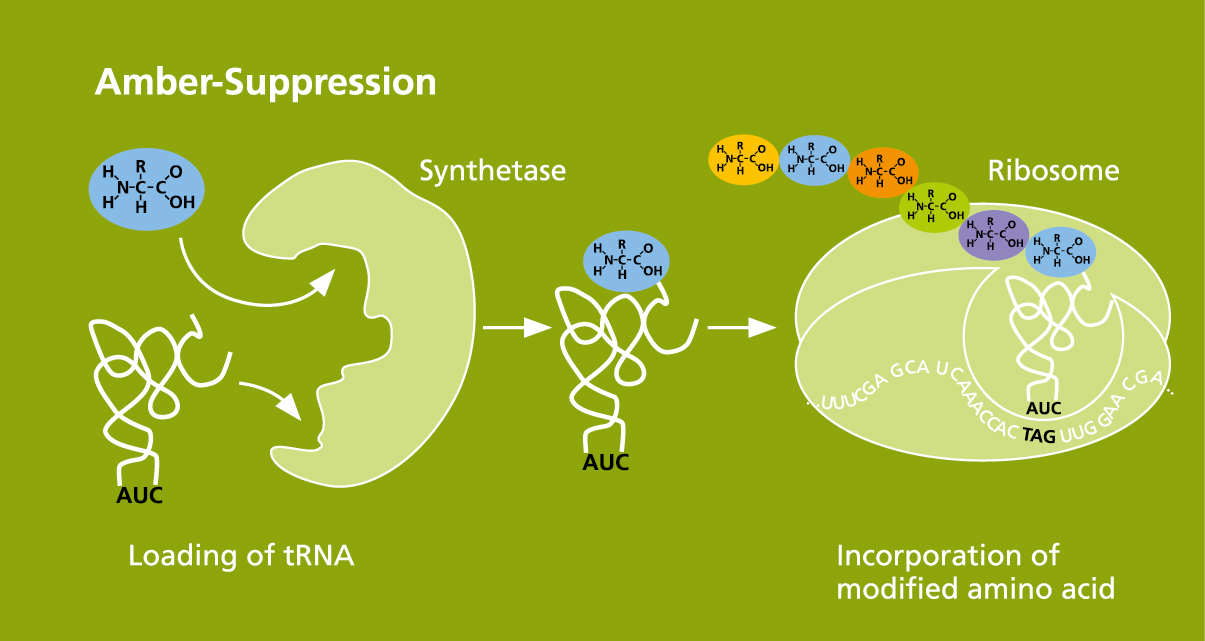
In addition to pure synthesis, the »Cell-free protein synthesis« research group offers a reliable method for introducing chemically modified amino acids into the cell-free synthesized proteins. The amber suppression method is used to introduce amino acids with a wide variety of reactive groups at site-specific positions. Furthermore, we use click chemistry to couple fluorescent dyes, sugar structures, PEGylations and biotinylations to the reactive groups of the introduced amino acids. Our method is well suited for the structural analysis and functional determination of membrane proteins, but also for the screening of novel ligands and therapeutics. In addition, proteins can be selectively modified to introduce new functions for therapeutic purposes.
Protein labeling and modification
Service offer
- Cell-free protein synthesis of difficult-to-produce proteins
- Embedding of chemically modified amino acids at integrated amber positions
- Coupling of fluorescent dyes, sugar residues, biotin and polymers (PEGs) to the target protein, both by copper-catalyzed and copper-free click reactions
- Design of DNA templates with matching amber codon position
- Tailor-made selection of the position and introduction of the amber codon into a specific gene sequence
- Evaluation of the efficiency of different amber codon positions to optimize the synthesis yield
- Analysis of the coupling efficiency
- Characterization of modified, cell-free synthesized proteins by microscopy and autoradiography
- Interaction and ligand binding studies
 Fraunhofer Institute for Cell Therapy and Immunology, Branch Bioanalytics and Bioprocesses IZI-BB
Fraunhofer Institute for Cell Therapy and Immunology, Branch Bioanalytics and Bioprocesses IZI-BB